Abstract
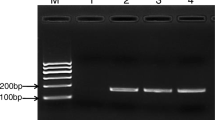
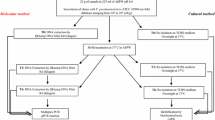
This study aimed to adopt MPN-PCR (most probable number-polymerase chain reaction) for rapid detection of the quantity of Vibrio parahaemolyticus in seafood. V. parahaemolyticus in seafood could be quantitated by MPN statistics according to PCR products. The sensitivity of MPN-PCR was 100 times higher than that of direct PCR. Of 225 seafood samples from Qingdao, 165 were positive for the presence of V. parahaemolyticus, with an MPN value of >719 per gram, and about 41.5% of samples were positive for tdh gene-possessing cells. Eighty muscle tissues from the 225 seafood samples were investigated by direct PCR and MPN-PCR, but no V. parahaemolyticus was detected. The MPN-PCR test could be completed in less than 16 h from the time of sample preparation. It was rapid, sensitive, and reliable for comprehensive detection and quick quantitative determination of V. parahaemolyticus in seafood and it revealed the potential risk of illness associated with their consumption.


Similar content being viewed by others
References
Alam MJ, Tomochika KI, Miyoshi SI (2002) Environmental investigation of potentially pathogenic Vibrio parahaemolyticus in the Seto-Inland Sea, Japan. FEMS Microbiol Lett 208:83–87
Bej AK, Patterson DP, Brasher CW, Vickery MCL, Jones DD, Kaysner CA (1999) Detection of total and hemolysin-producing Vibrio parahaemolyticus in shellfish using multiplex PCR amplification of tl, tdh and trh. J Microbiol Meth 36:215–225
Bilung LM, Radu S, Bahaman AR, Rahim RA, Napis S, Ling MWCV (2005) Detection of Vibrio parahaemolyticus in cockle (Anadara granosa) by PCR. FEMS Microbiol Lett 252:85–88
Hara-Kudo Y, Sugiyama K, Nishibuchi M et al (2003) Prevalence of pandemic thermostable direct hemolysin-producing Vibrio parahaemolyticus O3:K6 in Seafood and the coastal environment in Japan. Appl Environ Microb 69:3883–3891
Josephson KL, Gerba CP, Pepper RL (1993) Polymerase chain reaction detection of nonviable bacterial pathogens. Appl Environ Microb 59:3513–3515
Kaysner CA, Abeyta CJ, Scott RF, Krane MH, Wekell MM (1996) Enumeration of Vibrio species, including V. cholerae from samples of an oyster growing area, Grays Harbor, Washington. J Food Protect 53:300–302
Lee KK, Liu PC, Huang CY (2003) Vibrio parahaemolyticus infectious for both humans and edible mollusk abalone. Microbes Infect 5:481–485
Martín B, Jofré A, Garriga M, Hugas M, Aymerich T (2004) Quantification of Listeria monocytogenes in fermented sausages by MPN-PCR method. Lett Appl Microbiol 39:290–295
Miwa N, Nishio T, Arita Y, Kawamori F, Masuda T, Akiyama M (2003) Evaluation of MPN method combined with PCR procedure for detection and enumeration of Vibrio parahaemolyticus in seafood. Medline Abstr 44(6):289–293
Miwa N, Kashiwagi M, Kawamori F, Masuda T, Sano Y, Hiroi M, Kurashige H (2006) Levels of Vibrio parahaemolyticus and thermostable direct hemolysin gene-positive organisms in retail seafood determined by the most probable number-polymerase chain reaction (MPN-PCR) method. Medline Abstr 47(2):41–45
Picard C, Ponsonnet C, Paget E, Nesme X, Simonet P (1992) Detection and enumeration of bacteria in soil by direct DNA extraction and polymerase chain reaction. Appl Environ Microb 58:2717–2722
Scott E (1996) Foodborne disease and other hygienic issues in the home. J Appl Bacteriol 80:5–9
Tsai YL, Olson BH (1992) Detection of low number of cells in soils and sediments by polymerase chain reaction. Appl Environ Microb 58:754–757
Vesa M, Seppo N, Seppo K, Tuula P, Kristina L (1997) MPN-PCR-quantification method for staphylococcal enterotoxin c1 gene from fresh cheese. Int J Food Microbiol 36:135–143
Acknowledgments
This work was supported by Grant 30371108 from the National Natural Science Foundation of China and National High-Tech R&D Program 2003AA622070.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Luan, X., Chen, J., Liu, Y. et al. Rapid Quantitative Detection of Vibrio parahaemolyticus in Seafood by MPN-PCR. Curr Microbiol 57, 218–221 (2008). https://doi.org/10.1007/s00284-008-9177-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-008-9177-x